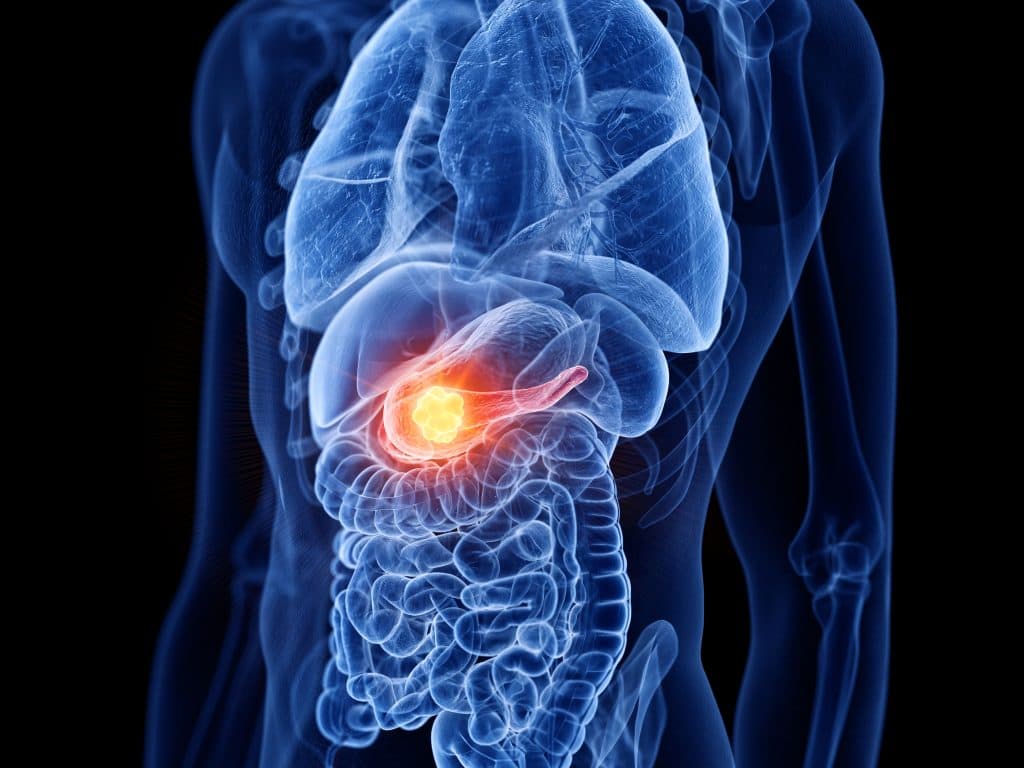
A bioengineer at the University of Texas at Dallas has developed synthetic enzymes that can control the behaviour of the signalling protein Vg1, which plays a key role in the development of muscle, bone and blood in vertebrate embryos.
The team of researchers is using a new approach, called the Synthetic Processing (SynPro) system, in zebrafish to study how the protein is formed.
By learning the molecular rules of signal formation in a developing animal, scientists aim to engineer mechanisms — such as giving cells new instructions — that could play a role in treating or preventing disease.
Dr P.C. Dave P. Dingal, assistant professor of bioengineering in the Erik Jonsson School of Engineering and Computer Science, and his colleagues published their research in Proceedings of the National Academy of Sciences.
Dingal said: “We’re interested in how synthetic enzymes might be used to control natural proteins, including disease-causing proteins.
“Our hope is to build biological circuits that, ultimately, we can introduce into cells and imbue them with new functions, like being able to detect cancer or resolve cellular disorders at the molecular level.”
The researcher said that zebrafish are ideal models for observing how signalling proteins are processed and secreted because zebrafish not only have approximately 70 per cent similarity with the human genome, but they also are small and easy to grow and image under the microscope.
The team studied interactions between the two proteins Vg1 and Nodal.
One of the questions the researchers investigated is why Vg1 remains inactive until it pairs with Nodal to form a larger protein complex called a heterodimer, which is secreted from cells and signals target embryonic cells to differentiate into specific tissues and organs.
Dingal said: “We discovered that there are proteins that act like chaperones that bind to Vg1 and force it to remain as an inactive monomer.
“In the presence of Nodal, however, the chaperones are released, and Nodal can then dimerize with Vg1.”
The scientists found that the act of pairing is not enough to activate Vg1 and Nodal. The Vg1 portion of the dimer must go through additional processing in other parts of the cell, including in the Golgi apparatus, where enzymes cut away, or cleave, unnecessary amino acids from the Vg1 section, much like a gardener prunes a rosebush.
To investigate the processing that Vg1 undergoes, the researchers developed a way to manipulate the protein.
Using a cleaving enzyme derived from a family of viruses, the scientist developed a synthetic enzyme that could be directed to cut specific amino acids from Vg1 in the zebrafish embryo.
They learned that Vg1-Nodal heterodimers do not need to undergo cleavage before they are released from the cell to bind with receptors on target cells.
Vg1, however, must undergo cleavage — while cleavage of Nodal is not required — to activate signalling on target cells.
Dingal will continue to study the proteins in the next phase of the project to determine, for example, the molecular rules that chaperone proteins use to control the composition of signalling complexes.
He recently received a $1.9 million (£1.5 million) grant from the National Institute of General Medical Sciences at the National Institutes of Health to continue his research.